 Author
Author  Correspondence author
Correspondence author
International Journal of Super Species Research, 2024, Vol. 14, No. 3
Received: 19 Mar., 2024 Accepted: 24 Apr., 2024 Published: 12 May, 2025
Escherichia coli ( E. coli) is a versatile bacterium that exists as a harmless commensal in the intestines of humans and animals but can transition into a pathogenic state, causing a variety of diseases. This study explores the mechanisms by which E. coli shifts from a commensal organism to a pathogen, focusing on four primary mechanisms of pathogenesis: quorum sensing, secretion systems, genetic variation, and virulence factors. Quorum sensing allows E. coli to regulate virulence factor expression in response to cell density, enhancing its pathogenic potential. The Type VI Secretion System (T6SS) is a nanomachine that injects toxic effector proteins into target cells, playing a crucial role in bacterial competition and virulence. Genetic variations, such as mutations in the FimH adhesin, can increase E. coli's ability to bind to host tissues, thereby enhancing its virulence. Additionally, E. coli can acquire virulence factors through horizontal gene transfer, enabling it to cause a wide range of diseases. Understanding these mechanisms is essential for developing new strategies to combat E. coli infections.
1 Introduction
Escherichia coli ( E. coli) is a versatile bacterium that predominantly resides as a commensal organism in the human gastrointestinal tract. As a commensal, E. coli plays a crucial role in maintaining gut health by contributing to the intestinal microbiota's balance and aiding in various physiological processes. The majority of E. coli strains in the human gut are harmless and coexist symbiotically with their host, contributing to the overall homeostasis of the intestinal environment (Tenaillon et al., 2010; Escribano-Vázquez et al., 2019; Martinson and Walk, 2020).
However, E. coli exhibits a dual nature, where certain strains can transition from being benign commensals to pathogenic organisms. Pathogenic E. coli strains are responsible for a wide range of diseases, both intestinal and extraintestinal. These pathogenic strains possess specific virulence factors that enable them to cause infections, ranging from mild gastroenteritis to severe conditions such as urinary tract infections, neonatal meningitis, and septicemia (Escribano-Vázquez et al., 2019; Denamur et al., 2020; Geurtsen et al., 2022). The pathogenicity of E. coli is often attributed to the acquisition of virulence genes through horizontal gene transfer, which can occur via plasmids, transposons, or bacteriophages (Geurtsen et al., 2022; Tawfick et al., 2022).
The objective of this study is to provide a comprehensive overview of E. coli's pathogenesis, focusing on the transition from commensal to pathogenic strains. This study will explore the genetic and environmental factors that contribute to the emergence of pathogenic E. coli, the mechanisms underlying their virulence, and the epidemiology of different E. coli pathotypes. By understanding the dual nature of E. coli, we aim to shed light on the complex interactions between commensal and pathogenic strains and the implications for human health. This study will also discuss current strategies for monitoring, preventing, and treating E. coli infections, emphasizing the need for continued research to address the challenges posed by this adaptable and often dangerous microorganism.
2 Biology and Ecology of E. coli
2.1 General characteristics of E. coli as a species
Escherichia coli (E. coli) is a Gram-negative, rod-shaped bacterium that belongs to the family Enterobacteriaceae. It is a facultative anaerobe, meaning it can survive in both aerobic and anaerobic environments. E. coli is predominantly found in the lower intestinal tract of warm-blooded animals, including humans, where it plays a significant role in the gut microbiome (Jang et al., 2017). The species exhibits remarkable genetic flexibility and adaptability, which allows it to thrive in various environments and acquire numerous antimicrobial resistance mechanisms (Ramos et al., 2020). This genetic plasticity is evident in the dynamic composition of its genome, which has undergone numerous lateral transfer events, contributing to its evolutionary success.
2.2 Commensal E. coli and its role in the human microbiome
As a commensal organism, E. coli is a natural inhabitant of the human gastrointestinal tract, where it coexists in a mutually beneficial relationship with its host. Approximately 90% of E. coli strains are commensals that rarely cause disease (Ramos et al., 2020). These commensal strains play a crucial role in maintaining gut health by contributing to the digestion of food, production of essential vitamins, and prevention of colonization by pathogenic bacteria (Jang et al., 2017). However, under certain conditions, such as inflammation, the balance of the gut microbiota can be disrupted, leading to an overgrowth of E. coli and potential pathogenicity (Winter et al., 2013; Kittana et al., 2018).
2.3 Mechanisms of E. coli colonization in the gut
E. coli colonizes the gut by adhering to the mucous layer of the colon, where it can utilize various mechanisms to establish and maintain its presence. One key factor in its colonization ability is its facultative anaerobic nature, which allows it to outcompete obligate anaerobes during episodes of gut inflammation by using host-derived nitrate for anaerobic respiration (Winter et al., 2013). Additionally, E. coli employs quorum sensing (QS) systems to regulate the expression of virulence factors and adapt to the surrounding environment. These QS systems include the orphan SdiA regulator, autoinducer-2 (AI-2), autoinducer-3 (AI-3), and indole, which facilitate communication and coordination among bacterial cells (Figure 1) (Mayer et al., 2023). The ability of E. coli to adapt to changing environmental conditions and its interactions with the host immune system further enhance its colonization and persistence in the gut (Kittana et al., 2018; Denamur et al., 2020).
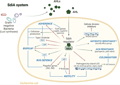 Figure 1 Schematic representation of the LuxR-type QS system in E. coli (Adopted from Mayer et al., 2023) Image caption: Although E. coli does not present a synthase that produces AHLs (green hexagon), it can sense these QS signals produced by other bacterial species. Through the orphan SdiA regulator, E. coli can induce a signalling to regulate the expression of several genes, including the ftsQAZ operon, in a cellular density-dependent manner. Effects on E. coli virulence-associated genes and phenotypes mediated by the LuxR solo regulator SdiA are shown. Green arrows indicate positive regulation, red lines indicate gene or phenotype inhibition, and blue lines indicate a link between different phenotypes. *Biofilm regulation by SdiA is a controversial point in the literature as explained below. **Although SdiA plays a role in drug resistance regulation, the effect of AHLs on this phenotype (dotted line) needs to be confirmed (Adopted from Mayer et al., 2023) |
3 Pathogenic Strains of E. coli
3.1 Classification of pathogenic E. coli strains
Pathogenic Escherichia coli ( E. coli) strains are classified into several pathotypes based on their virulence factors and the diseases they cause. The major pathotypes include Enteropathogenic E. coli (EPEC), Enterohemorrhagic E. coli (EHEC), and Uropathogenic E. coli (UPEC). EPEC is a significant cause of infant diarrhea in developing countries and is characterized by its ability to form attaching and effacing lesions on the intestinal epithelium (Hazen et al., 2017). EHEC, including the notorious O157:H7 strain, is associated with severe conditions such as hemorrhagic colitis and hemolytic uremic syndrome, primarily due to the production of Shiga toxins. UPEC is a leading cause of urinary tract infections (UTIs) and is distinguished by its ability to colonize the urinary tract and evade host defenses (Russo and Johnson, 2000).
3.2 Genetic determinants of virulence in pathogenic strains
The virulence of pathogenic E. coli strains is largely determined by the presence of specific genetic elements, including pathogenicity islands, plasmids, and bacteriophages, which encode various virulence factors. For instance, EPEC strains possess the locus of enterocyte effacement (LEE) pathogenicity island, which encodes a type III secretion system (T3SS) and effector proteins that disrupt host cell processes (Hazen et al., 2017; Donnenberg and Whittam, 2001). EHEC strains carry genes for Shiga toxins (stx1 and stx2) on lambdoid prophages, which are critical for their pathogenicity (Donnenberg and Whittam, 2001). UPEC strains harbor a variety of virulence genes, such as those encoding adhesins (e.g., fimH), toxins, and iron acquisition systems, often located on plasmids and pathogenicity islands (Khairy et al., 2019). The acquisition of these genetic elements through horizontal gene transfer contributes to the diversity and adaptability of pathogenic E. colistrains.
3.3 Comparison of pathogenic E. coli with commensal strains
Pathogenic E. coli strains differ significantly from commensal strains in their genetic makeup and pathogenic potential. While commensal E. coli strains are typically harmless and form part of the normal gut flora, pathogenic strains possess additional virulence factors that enable them to cause disease. Commensal strains generally lack the specialized virulence traits found in pathogenic strains, such as the T3SS in EPEC, Shiga toxins in EHEC, and the array of adhesins and toxins in UPEC (Russo and Johnson, 2000). Moreover, pathogenic strains often exhibit greater genetic plasticity, with frequent acquisition and loss of virulence genes through horizontal gene transfer, which is less common in commensal strains (Halaji et al., 2022). This genetic fluidity allows pathogenic E. coli to adapt to different environments and hosts, enhancing their ability to cause a wide range of diseases (Khairy et al., 2019).
4 Mechanisms of Pathogenesis
4.1 Adhesion and invasion mechanisms
Pathogenic Escherichia coli ( E. coli) employ a variety of adhesion and invasion mechanisms to establish infection within the host. Uropathogenic E. coli (UPEC), responsible for urinary tract infections, utilize adhesive organelles such as type 1, P, and S pili, along with Dr adhesins, to attach to and invade uroepithelial cells. This invasion allows the bacteria to replicate within host cells, providing a survival advantage by evading immune detection and clearance (Mulvey, 2002). Similarly, Shiga toxin-producing E. coli (STEC) strains use the intimin adhesin to mediate attachment to the intestinal epithelium, a critical step for colonization and subsequent toxin production (Farfán and Torres, 2011). Additionally, Locus of Enterocyte Effacement (LEE)-negative STEC strains utilize the tia gene, which is involved in the invasion of epithelial cell monolayers, further highlighting the diverse strategies employed by E. coli to adhere to and invade host tissues (Bondí et al., 2017).
4.2 Toxin production and its role in disease manifestation
Toxin production is a key virulence factor in the pathogenesis of E. coli infections. Shiga toxins (Stxs) produced by STEC are responsible for severe conditions such as hemorrhagic colitis and hemolytic uremic syndrome (HUS). These toxins disrupt protein synthesis in host cells, leading to cell death and tissue damage (Farfán and Torres, 2011). The role of toxins in disease manifestation is further exemplified by the Secreted Autotransporter Toxin (Sat), which not only exhibits cytotoxic effects but also modulates the host immune response by cleaving complement proteins, thereby aiding in immune evasion and contributing to the severity of bloodstream infections and sepsis (Freire et al., 2022). The production of these toxins underscores the pathogenic potential of E. coli and their ability to cause significant morbidity and mortality.
4.3 Immune evasion strategies employed by pathogenic E. coli
Pathogenic E. coli have evolved multiple strategies to evade the host immune system, ensuring their survival and persistence within the host. One such strategy involves the production of serine proteases like Pic, which cleave complement proteins, thereby reducing complement activation and protecting the bacteria from complement-mediated killing (Abreu et al., 2015). Similarly, the Sat toxin has been shown to cleave various complement proteins, further aiding in immune evasion (Figure 2) (Freire et al., 2022). These proteolytic activities not only help E. coli to evade the innate immune response but also enhance their ability to cause severe infections. Additionally, the ability of UPEC to invade and replicate within host cells allows them to avoid detection and clearance by both innate and adaptive immune mechanisms, further illustrating the sophisticated immune evasion strategies employed by pathogenic E. coli.
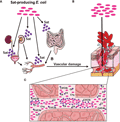 Figure 2 Hypothetical model of Sat enrollment in BSI and sepsis pathogenesis (Adopted from Freire et al., 2022) Image caption: A: Sat-producing E. coli present in the urinary tract or in the intestinal tract secretes Sat, causing cellular damage in the bladder, kidneys, or intestine. This damage allows the bacteria to move forward to the blood vessels, whose endothelial cells are also susceptible to Sat cytotoxic action; B: Vascular damage caused by Sat allows E. coli to access the bloodstream; C: Sat secretion in the bloodstream protects E. coli from complement killing, by direct cleavage of C2, C3, C3b, C4, C4b, C5, C6, C7, C8 and C9. This evasion mechanism allows Sat-producing E. coli to multiply and disseminate in the bloodstream and consequently reach other organs such as the spleen and the liver, where new sites of infection can be established and a magnification of the immune response can occur, facilitating the progression of the disease to sepsis (Adopted from Freire et al., 2022) |
5 Environmental and Host Factors Influencing Pathogenicity
5.1 Influence of the host’s immune system on E. coli pathogenicity
The host's immune system plays a crucial role in influencing the pathogenicity of Escherichia coli. The dynamic relationship between commensal bacteria and the host's immune response is pivotal in determining the outcome of infections. Commensal bacteria can modulate the immune system by providing tonic stimulation that induces the basal expression of factors required for host defense, thereby enhancing the host's ability to combat invasive pathogens (Sarowska et al., 2019). However, dysbiosis, or the imbalance in the microbial community, can lead to improper training of the immune system, resulting in increased susceptibility to immune-mediated diseases such as inflammatory bowel disease and multiple sclerosis (Petersen and Round, 2014). This improper immune response can create an environment conducive to the emergence of pathogenic E. coli strains, which can exploit the weakened immune defenses to establish infections.
5.2 Role of environmental factors in the emergence of pathogenic strains
Environmental factors, particularly antibiotic use and diet, significantly contribute to the emergence of pathogenic E. coli strains. The use of antibiotics in food-producing animals has been implicated in the development of antimicrobial resistance among commensal E. coli populations. This resistance can be transferred to pathogenic strains, leading to the emergence of multidrug-resistant bacteria (Ramos et al., 2020). The genetic flexibility of E. coli allows it to acquire resistance mechanisms through horizontal gene transfer, further complicating the treatment of infections (Navarro-García et al., 2019). Additionally, dietary factors can influence the gut microbiota composition, potentially promoting the growth of pathogenic strains over commensal ones. The selective pressure exerted by antibiotics and dietary changes can thus drive the evolution of E. coli from a benign commensal to a harmful pathogen.
5.3 Cross-species transmission and zoonotic potential of pathogenic E. coli
Pathogenic E. coli strains have a significant zoonotic potential, with the ability to cross species barriers and infect both humans and animals. The transmission of E. coli from food-producing animals to humans is a major public health concern, particularly with strains that produce extended-spectrum β-lactamases (ESBLs), which confer resistance to critical antibiotics (Ramos et al., 2020). The genetic similarity between E. coli strains isolated from humans, animals, and food suggests that cross-species transmission is a common occurrence (Sarowska et al., 2019). This zoonotic potential is facilitated by the acquisition of virulence factors through horizontal gene transfer, enabling E. coli to adapt to different hosts and environments (Figure 3) (Desvaux et al., 2020). Understanding the mechanisms underlying this transmission is essential for developing strategies to prevent the spread of pathogenic E. coli and protect public health.
 Figure 3 Structure and motility functions of PAIs (Adopted from Desvaux et al., 2020) Image caption: A: Insertion and excision processes of PAIs. PAIs are chromosomal fragments of pathogenic bacteria that encode biological functions involved in virulence. Their insertion in the chromosome is due to the presence of att sites at a chromosomal acceptor site (attB) and in the episomal PAI (attP). They are recognized by integrases, which catalyze a recombination of att sites. It results in the insertion of the episomal element at the attB site and the formation of direct repeated sequences (DRS) also named attL (left DRS) and attR (right DRS) in the ends of the inserted PAI. The excision of the PAI results from recombination between the direct repeats attL and attR. Catalyzed by integrases and recombination directionality factors (RDFs) also called excisionases, it generates an episomal element that contains one of the att sites (attP), while the other att site remains in the chromosome (attB); B: Horizontal transfer of PAIs via conjugative plasmids, ICEs, and phages harboring att sites. Episomal PAIs can be inserted at att sites in conjugative plasmids, ICEs and phages as described above and then transferred into a bacterial recipient via conjugation for ICE-type and plasmid-type navettes or via transduction for phage-type navettes (Adopted from Desvaux et al., 2020) |
6 Case Study
6.1 Background and epidemiology of the outbreak
In May 2011, Germany experienced one of the largest outbreaks of entero-hemorrhagic Escherichia coli (EHEC) ever reported, primarily affecting the city of Hamburg. The outbreak was caused by a highly virulent strain of E. coli O104:H4, which led to 3 128 cases of acute gastroenteritis, 782 cases of hemolytic-uremic syndrome (HUS), and 46 deaths. The initial epidemiological signal was detected when local health authorities identified a cluster of HUS cases, by which time 99 EHEC and 48 HUS patients had already experienced symptoms (Mellmann et al., 2011). Middle-aged women were found to be at the highest risk of contracting the infection. The outbreak strain was notable for its high virulence and resistance, which contributed to the severity and rapid spread of the disease (Navarro-García, 2014).
6.2 Molecular characterization of the EHEC strain involved
The E. coli O104:H4 strain responsible for the outbreak was a hybrid of enteroaggregative E. coli (EAEC) and Enterohemorrhagic E. coli (EHEC), combining virulence factors from both pathotypes. This strain contained a prophage encoding Shiga toxin 2, a characteristic feature of EHEC, and a set of virulence-associated genes regulated by the AggR transcription factor, typical of EAEC (Mellmann et al., 2011). Whole genome sequencing revealed that the outbreak strain was closely related to the EAEC strain 55989 but had acquired additional virulence and antibiotic-resistance factors through horizontal genetic exchange. The strain's ability to adhere to intestinal epithelial cells and produce Shiga toxin contributed to its high pathogenicity and the severe clinical outcomes observed during the outbreak (Rasko et al., 2011).
6.3 Public health response and lessons learned from the outbreak
The public health response to the 2011 EHEC outbreak involved rapid epidemiological investigations, enhanced surveillance, and inter-agency collaboration. The outbreak highlighted the importance of timely case notification and the need for robust public health infrastructure to manage such crises (Tahden et al., 2016). Research conducted in the aftermath of the outbreak emphasized the necessity for improved methods in surveillance, diagnosis, and outbreak investigation, as well as the critical role of public communication in managing public health emergencies (Köckerling et al., 2019). The outbreak also underscored the need for international cooperation, as most research publications were conducted by multinational teams. Lessons learned from this outbreak have informed public health strategies and preparedness plans for future outbreaks of similar nature.
7 Clinical manifestations of E. coli infections
7.1 Common diseases caused by pathogenic E. coli
Pathogenic Escherichia coli ( E. coli) strains are responsible for a variety of diseases, with urinary tract infections (UTIs) being among the most prevalent. Uropathogenic E. coli (UPEC) is the primary causative agent of both uncomplicated and complicated UTIs, which can range from cystitis (bladder infection) to pyelonephritis (kidney infection) and even urosepsis (Mobley et al., 2009; Alteri and Mobley, 2015; Zhou et al., 2023). UPEC strains possess unique virulence factors, such as fimbriae and toxins, which enable them to adhere to and invade the urinary tract epithelium, leading to infection (Terlizzi et al., 2017). Additionally, E. coli is a common cause of gastroenteritis, particularly through strains like Enterotoxigenic E. coli (ETEC) and Enterohemorrhagic E. coli (EHEC), which can cause severe diarrheal diseases (Alteri and Mobley, 2015).
7.2 Complications associated with E. coli infections
Complications arising from E. coli infections can be severe and life-threatening. One notable complication is hemolytic uremic syndrome (HUS), which is often associated with EHEC infections. HUS is characterized by hemolytic anemia, thrombocytopenia, and acute kidney injury, and it primarily affects children (Alteri and Mobley, 2015). In the context of UTIs, complications can include pyelonephritis, which may lead to renal scarring and chronic kidney disease, and urosepsis, a severe systemic infection that can result in septic shock and multi-organ failure (Terlizzi et al., 2017). The emergence of multidrug-resistant UPEC strains further complicates treatment and increases the risk of severe outcomes (Totsika et al., 2012; Zhou et al., 2023).
7.3 Diagnostic approaches for identifying E. coli infections
Accurate and timely diagnosis of E. coli infections is crucial for effective treatment. Diagnostic approaches for UTIs typically involve urine culture to identify the presence of UPEC and determine antibiotic susceptibility (Zhou et al., 2023; Totsika et al., 2012). Molecular techniques, such as polymerase chain reaction (PCR), can also be used to detect specific virulence genes associated with pathogenic E. coli strains (Rezatofighi et al., 2021). For gastroenteritis caused by E. coli, stool samples are analyzed to identify the presence of pathogenic strains, often using culture methods and PCR to detect specific toxins or virulence factors. Advances in genomic and proteomic technologies are enhancing our ability to rapidly and accurately diagnose E. coli infections, which is essential for managing and controlling these diseases (Biggel et al., 2020; Mann et al., 2017).
8 Antibiotic Resistance in Pathogenic E. coli
8.1 Mechanisms of antibiotic resistance in E. coli
Antibiotic resistance in Escherichia coli ( E. coli) is a multifaceted problem driven by various genetic and biochemical mechanisms. E. coli can acquire resistance through horizontal gene transfer, which involves the exchange of resistance genes via plasmids, transposons, and integrons (Szmolka and Nagy, 2013; Blair et al., 2014). These genetic elements can carry multiple resistance genes, leading to multidrug-resistant (MDR) strains. Additionally, E. coli can develop resistance through mutations in its genome that alter drug targets, enhance efflux pump activity, or modify membrane permeability. For instance, efflux pumps can actively expel antibiotics from the bacterial cell, reducing drug accumulation and efficacy (Gauba and Rahman, 2023). Enzymatic degradation or modification of antibiotics, such as beta-lactamase production, is another common resistance mechanism (Džidić et al., 2008). Understanding these mechanisms is crucial for developing strategies to combat antibiotic resistance in E. coli.
8.2 Global trends in antibiotic-resistant E. coli strains
The prevalence of antibiotic-resistant E. coli strains is a growing global health concern. Studies have shown high levels of resistance in E. coli isolates from various sources, including humans, animals, food, and the environment. For example, a systematic study and meta-analysis revealed significant resistance rates to commonly used antibiotics such as ampicillin, cefotaxime, and ciprofloxacin in community settings, particularly in low- and middle-income countries (LMICs) (Nji et al., 2021). Another study highlighted the widespread presence of MDR E. coli in food-producing animals, which can act as reservoirs for resistance genes that may transfer to human pathogens (Szmolka and Nagy, 2013). The increasing resistance trends underscore the need for continuous monitoring and global cooperation to address this public health threat (Pormohammad et al., 2019; Urban-Chmiel et al., 2022).
8.3 Implications for treatment and public health
The rise of antibiotic-resistant E. coli strains has profound implications for treatment and public health. The effectiveness of standard antibiotic therapies is diminishing, leading to higher morbidity and mortality rates associated with E. coli infections (Urban-Chmiel et al., 2022). This situation necessitates the development of novel therapeutic strategies and the prudent use of existing antibiotics to slow the spread of resistance (Munita and Arias, 2016). Public health policies must focus on surveillance, infection control, and the promotion of antibiotic stewardship programs to mitigate the impact of antibiotic resistance (Nji et al., 2021; Pormohammad et al., 2019). Additionally, there is a critical need for research into alternative treatments, such as bacteriophage therapy and the use of narrow-spectrum antibiotics, to provide effective options against resistant E. coli strains (Gauba and Rahman, 2023). Addressing antibiotic resistance in E. coli is essential to safeguarding global health and ensuring the continued efficacy of antimicrobial therapies.
9 Prevention and Control Strategies
9.1 Hygiene and sanitation measures to prevent E. coli infections
Hygiene and sanitation are fundamental in preventing E. coli infections. Proper handwashing, especially after using the restroom and before handling food, is crucial. Ensuring clean water supply and proper sewage disposal can significantly reduce the risk of E. coli contamination. Additionally, food safety practices such as thoroughly cooking meat, washing fruits and vegetables, and avoiding cross-contamination in the kitchen are essential preventive measures. These practices help in minimizing the transmission of E. coli from contaminated sources to humans.
9.2 Vaccination and novel therapeutic approaches
Vaccination represents a promising strategy to prevent E. coli infections. Several vaccine candidates are under investigation, including the ExPEC4V, a bioconjugate vaccine targeting extraintestinal pathogenic E. coli (ExPEC). This vaccine has shown to be well-tolerated and capable of eliciting robust antibody responses in clinical trials, although further studies are needed to confirm its efficacy in reducing infection rates (Huttner et al., 2017; Magistro and Stief, 2019; Naili et al., 2023). Another approach involves the use of live attenuated bacterial vaccines, which have demonstrated protective immunity in animal models, particularly in immunocompromised hosts (Wang and Wan, 2023). Additionally, the identification of conserved antigens such as OmpA and SslE offers potential for developing broad-spectrum vaccines that could protect against multiple E. coli pathotypes.
9.3 Public health policies and their role in controlling the spread of pathogenic E. coli
Public health policies play a critical role in controlling the spread of pathogenic E. coli. These policies include surveillance programs to monitor E. coli outbreaks, guidelines for antibiotic use to prevent resistance, and public education campaigns on hygiene practices. The implementation of antimicrobial stewardship programs is essential to manage the use of antibiotics and reduce the emergence of resistant E. coli strains (Magistro and Stief, 2019; Mayer et al., 2023). Furthermore, policies promoting research and development of new vaccines and treatments are vital to stay ahead of evolving E. coli strains and ensure public health safety.
10 Concluding Remarks
Escherichia coli ( E. coli) is a versatile bacterium that exists both as a harmless commensal organism in the intestines of humans and animals and as a formidable pathogen responsible for a variety of infections. The transition from commensalism to pathogenicity in E. coli is driven by several mechanisms, including the acquisition of virulence genes through horizontal gene transfer and the regulation of these genes via quorum sensing systems. The genetic diversity among E. coli strains is substantial, with significant differences in virulence potential and antimicrobial resistance observed between commensal and pathogenic strains. The emergence of multidrug-resistant strains, particularly those producing extended-spectrum β-lactamases (ESBL), poses a significant public health challenge. Furthermore, the ability of commensal E. coli to act as a reservoir for virulence genes and antibiotic resistance highlights the complex interplay between commensal and pathogenic states.
The findings underscore the need for a deeper understanding of the genetic and environmental factors that drive the transition from commensalism to pathogenicity in E. coli. Future research should focus on elucidating the specific genetic determinants and regulatory networks involved in this process. Additionally, the role of the gut microbiota in providing colonization resistance against pathogenic E. coli strains warrants further investigation, as this could inform the development of probiotic and prebiotic therapies aimed at enhancing gut health and preventing infections. Public health strategies should prioritize the monitoring of antimicrobial resistance patterns in both commensal and pathogenic E. coli populations to inform effective antibiotic stewardship programs.
To improve the management and prevention of E. coli infections, several recommendations can be made. First, enhancing surveillance systems to track the prevalence and spread of virulent and antibiotic-resistant E. coli strains is crucial. Second, promoting the prudent use of antibiotics in both human medicine and animal husbandry can help mitigate the emergence of resistant strains. Third, developing targeted vaccines against the most common and virulent E. coli pathotypes could provide a proactive approach to preventing infections. Finally, public health initiatives should focus on educating the public about proper hygiene practices and safe food handling to reduce the risk of E. coli transmission. By implementing these strategies, we can better manage and prevent E. coli infections, ultimately improving public health outcomes.
![]() Acknowledgments
Acknowledgments
The EcoEvo Publisher sincerely thanks the two anonymous peer reviewers for their review of this manuscript and for providing constructive revision suggestions.
Conflict of Interest Disclosure
The authors affirm that this research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
Abreu A., Fraga T., Martinez A., Kondo M., Juliano M., Juliano L., Navarro-García F., Isaac L., Barbosa A., and Elias W., 2015, The serine protease Pic from Enteroaggregative Escherichia coli mediates immune evasion by the direct cleavage of complement proteins, J. Infect. Dis., 212(1): 106-115.
https://doi.org/10.1093/infdis/jiv013
Alteri C., and Mobley H., 2015, Metabolism and fitness of urinary tract pathogens, Microbiol. Spectr., 3: 3.
https://doi.org/10.1128/microbiolspec.MBP-0016-2015
Biggel M., Xavier B., Johnson J., Nielsen K., Frimodt-Møller N., Matheeussen V., Goossens H., Moons P., and Puyvelde S. V., 2020, Horizontally acquired papGII-containing pathogenicity islands underlie the emergence of invasive uropathogenic Escherichia coli lineages, Nat. Commun., 11(1): 5968.
https://doi.org/10.1038/s41467-020-19714-9
Blair J., Webber M., Baylay A., Ogbolu D., and Piddock L., 2014, Molecular mechanisms of antibiotic resistance, Nat. Rev. Microbiol., 13: 42-51.
https://doi.org/10.1038/nrmicro3380
Bondí R., Chiani P., Michelacci V., Minelli F., Caprioli A., and Morabito S., 2017, The gene tia, harbored by the subtilase-encoding pathogenicity island, is involved in the ability of locus of enterocyte effacement-negative Shiga toxin-producing Escherichia coli strains to invade monolayers of epithelial cells, Infect. Immun., 85(12): e00613-17.
https://doi.org/10.1128/IAI.00613-17
Denamur E., Clermont O., Bonacorsi S., and Gordon D., 2020, The population genetics of pathogenic Escherichia coli, Nat. Rev. Microbiol., 19: 37-54.
https://doi.org/10.1038/s41579-020-0416-x
Desvaux M., Dalmasso G., Beyrouthy R., Barnich N., Delmas J., and Bonnet R., 2020, Pathogenicity factors of genomic islands in intestinal and extraintestinal Escherichia coli, Front. Microbiol., 11: 2065.
https://doi.org/10.3389/fmicb.2020.02065
Donnenberg M., and Whittam T., 2001, Pathogenesis and evolution of virulence in enteropathogenic and enterohemorrhagic Escherichia coli, J. Clin. Invest., 107(5): 539-548.
https://doi.org/10.1172/JCI12404
Džidić S., Šušković J., and Kos B., 2008, Antibiotic resistance mechanisms in bacteria: biochemical and genetic aspects, Food Technol. Biotechnol., 46: 11-21.
Escribano-Vázquez U., Ramírez-Rico G., and Godoy-Hernández G., 2020, Host–parasitoid interactions and community structure of braconid wasps (Hymenoptera: Braconidae) in tropical agroecosystems, Journal of Insect Conservation, 24: 1037-1048.
https://doi.org/10.1007/s10841-020-00263-3
Farias-Silva F., and Santos M., 2019, Ecological networks of parasitoid–host relationships: Structure and implications for biological control, Ecological Entomology, 44: 394-406.
https://doi.org/10.1111/een.12710
Fathipour Y., and Maleknia B., 2016, Population dynamics and biological control of aphids by parasitoids and predators: A review, Journal of Entomological Society of Iran, 36: 13-33.
Fernández-Arhex V., and Corley J. C., 2003, The functional response of parasitoids and its implications for biological control, Biocontrol Science and Technology, 13: 403-413.
https://doi.org/10.1080/0958315031000104523
Giron D., Pincebourde S., Casas J., and Mandon N., 2022, Physiological adaptations of parasitoid wasps: An integrative overview, Annual Review of Entomology, 67: 195-216.
https://doi.org/10.1146/annurev-ento-011021-044159
Gross P., 1993, Insect behavioral and morphological defenses against parasitoids, Annual Review of Entomology, 38: 251-273.
https://doi.org/10.1146/annurev.en.38.010193.001343
Hafer-Hahmann N., 2019, Parasitoid manipulation of host behaviour: From evolutionary ecology to mechanisms, Behavioural Processes, 159: 60-66.
https://doi.org/10.1016/j.beproc.2018.12.007
Harvey J. A., and Malcicka M., 2016, Nutritional integration between insect hosts and koinobiont parasitoids in an evolutionary framework, Entomologia Experimentalis et Applicata, 159: 181-188.
https://doi.org/10.1111/eea.12433
Henry L. M., Gillespie D. R., and Roitberg B. D., 2005, Does mother really know best? Oviposition preference reduces reproductive performance in the generalist parasitoid Aphidius ervi, Entomologia Experimentalis et Applicata, 116: 167-174.
https://doi.org/10.1111/j.1570-7458.2005.00317.x
Hopper K. R., 1999, Risk‐spreading and bet‐hedging in insect population biology, Annual Review of Entomology, 44: 535-560.
https://doi.org/10.1146/annurev.ento.44.1.535
Hudson P. J., Dobson A. P., and Lafferty K. D., 2006, Is a healthy ecosystem one that is rich in parasites? Trends in Ecology & Evolution, 21: 381-385.
https://doi.org/10.1016/j.tree.2006.04.007
Jervis M. A., Ellers J., and Harvey J. A., 2008, Resource acquisition, allocation, and utilization in parasitoid reproductive strategies, Annual Review of Entomology, 53: 361-385.
https://doi.org/10.1146/annurev.ento.53.103106.093433
Jones O. R., and Purvis A., 2020, Comparative methods in evolutionary biology: A practical approach for testing adaptive hypotheses, Biological Reviews, 95: 1499-1524.
https://doi.org/10.1111/brv.12620
Korenko S., and Pekár S., 2011, Is there an evolutionary trade-off between parasitism and predation? A case study of a spider parasitoid Zatypota percontatoria, Evolutionary Ecology, 25: 757-769.
https://doi.org/10.1007/s10682-010-9434-7
Kraaijeveld A.R., and Godfray H. C.J., 1999, Geographic patterns in the evolution of resistance and virulence in Drosophila–parasitoid interactions, American Naturalist, 153: S61-S74.
https://doi.org/10.1086/303212
Laurent R., Frago E., and Gagic V., 2021, Trophic interactions under climate change: Effects on host–parasitoid systems and biological control, Biological Control, 158: 104600.
https://doi.org/10.1016/j.biocontrol.2021.104600
Lins J. C., Oliveira V. M., and Queiroz J. M., 2019, Morphological adaptations of parasitoid wasps to host exploitation, Zoologischer Anzeiger, 281: 21-31.
https://doi.org/10.1016/j.jcz.2019.04.004
Moreau S. J. M., and Asgari S., 2015, Venom proteins from parasitoid wasps and their biological functions, Toxins, 7: 2385-2412.
https://doi.org/10.3390/toxins7072385
Murdock C. C., Paaijmans K. P., Cox-Foster D., Read A. F., and Thomas M. B., 2012, Rethinking vector immunology: The role of environmental temperature in shaping resistance, Nature Reviews Microbiology, 10: 869-876.
https://doi.org/10.1038/nrmicro2900
Naeem A., 2020, Integrative approaches to parasitoid biodiversity and ecosystem stability, Biodiversity and Conservation, 29: 1071-1086.
https://doi.org/10.1007/s10531-019-01893-0
Ramos S., and Castaño-Meneses G., 2021, Functional diversity and trophic roles of parasitoid wasps in Neotropical ecosystems, Ecological Entomology, 46: 411-422.
https://doi.org/10.1111/een.12989
Rasko D.A., Webster D.R., Sahl J.W., Bashir A., Boisen N., Scheutz F., Paxinos E.E., Sebra R., Chin C.S., Iliopoulos D., Klammer A., Peluso P., Lee L., Kislyuk A.O., Bullard J., Kasarskis A., Wang S., Eid J., Rank D., Redman J.C., Steyert S.R., Frimodt-Møller J., Struve C., Petersen A.M., Krogfelt K.A., Nataro J.P., Schadt E.E., and Waldor M.K., 2011, Origins of the E. coli strain causing an outbreak of hemolytic-uremic syndrome in Germany, The New England Journal of Medicine, 365(8): 709-717.
https://doi.org/10.1056/NEJMoa1106920
Rezatofighi S.E., Mirzarazi M., and Salehi M., 2021, Virulence genes and phylogenetic groups of uropathogenic Escherichia coli isolates from patients with urinary tract infection and uninfected control subjects: a case-control study, BMC Infectious Diseases, 21(1): 361.
https://doi.org/10.1186/s12879-021-06036-4
Russo T.A., and Johnson J.R., 2000, Proposal for a new inclusive designation for extraintestinal pathogenic isolates of Escherichia coli: ExPEC, The Journal of Infectious Diseases, 181(5): 1753-1754.
https://doi.org/10.1086/315418
Sarowska J., Futoma-Kołoch B., Jama-Kmiecik A., Frej-Mądrzak M., Książczyk M., Bugla-Płoskońska G., and Choroszy-Król I., 2019, Virulence factors, prevalence and potential transmission of extraintestinal pathogenic Escherichia coli isolated from different sources: recent reports, Gut Pathogens, 11(1): 10.
https://doi.org/10.1186/s13099-019-0290-0
Szmolka A., and Nagy B., 2013, Multidrug resistant commensal Escherichia coli in animals and its impact for public health, Frontiers in Microbiology, 4: 258.
https://doi.org/10.3389/fmicb.2013.00258
Tahden M., Manitz J., Baumgardt K., Fell G., Kneib T., and Hegasy G., 2016, Epidemiological and ecological characterization of the EHEC O104:H4 outbreak in Hamburg, Germany, 2011, PLoS ONE, 11(10): e0164508.
https://doi.org/10.1371/journal.pone.0164508
Tawfick M., Elshamy A., Mohamed K., and Menofy N., 2022, Gut commensal Escherichia coli, a high-risk reservoir of transferable plasmid-mediated antimicrobial resistance traits, Infection and Drug Resistance, 15: 1077-1091.
https://doi.org/10.2147/IDR.S354884
Tenaillon O., Skurnik D., Picard B., and Denamur E., 2010, The population genetics of commensal Escherichia coli, Nature Reviews Microbiology, 8: 207-217.
https://doi.org/10.1038/nrmicro2298
Terlizzi M.E., Gribaudo G., and Maffei M.E., 2017, Uropathogenic Escherichia coli (UPEC) infections: virulence factors, bladder responses, antibiotic, and non-antibiotic antimicrobial strategies, Frontiers in Microbiology, 8: 1566.
https://doi.org/10.3389/fmicb.2017.01566
Totsika M., Moriel D.G., Idris A., Rogers B.A., Wurpel D.J., Phan M.D., Paterson D.L., and Schembri M.A., 2012, Uropathogenic Escherichia coli-mediated urinary tract infection, Current Drug Targets, 13(11): 1386-1399.
https://doi.org/10.2174/138945012803530206
Urban-Chmiel R., Marek A., Stępień-Pyśniak D., Wieczorek K., Dec M., Nowaczek A., and Osek J., 2022, Antibiotic resistance in bacteria—a review, Antibiotics, 11(8): 1079.
https://doi.org/10.3390/antibiotics11081079
Wang S., and Wan F., 2023, Oral vaccination with live attenuated pathogenic bacteria protects immunocompromised hosts from lethal colitis, The Journal of Immunology, 210(Supplement_1): 253.08.
https://doi.org/10.4049/jimmunol.210.supp.253.08
Winter S.E., Winter M.G., Xavier M.N., Thiennimitr P., Poon V., Keestra A.M., Laughlin R.C., Gomez G., Wu J., Lawhon S.D., Popova I.E., Parikh S.J., Adams L.G., Tsolis R.M., Stewart V.J., and Bäumler A.J., 2013, Host-derived nitrate boosts growth of E. coli in the inflamed gut, Science, 339(6120): 708-711.
https://doi.org/10.1126/science.1232467
Wu X.R., Sun T.T., and Medina J.J., 1996, In vitro binding of type 1-fimbriated Escherichia coli to uroplakins Ia and Ib: relation to urinary tract infections, Proceedings of the National Academy of Sciences of the United States of America, 93(18): 9630-9635.
https://doi.org/10.1073/pnas.93.18.9630
Zhou Y., Zhou Z., Zheng L., Gong Z., Li Y., Jin Y., Huang Y., and Chi M., 2023, Urinary tract infections caused by uropathogenic Escherichia coli: mechanisms of infection and treatment options, International Journal of Molecular Sciences, 24(13): 10537.
https://doi.org/10.3390/ijms241310537

. HTML
Associated material
. Readers' comments
Other articles by authors
. Shusheng Liu
. Fumin Gao
Related articles
. Quorum sensing
. Type VI secretion
. System genetic
. Variation virulence factors
. Pathogenesis
Tools
. Post a comment


